Sentinel-3#
Let’s use EOReader to open and read some Sentinel-3 bands and see the specificities of OLCI and SLSTR constellations.
Note that Sentinel-3 processes have stopped to use SNAP since the 0.8.0 version.
Some minor discrepancies might occur both in the geocoding (especially OLCI) and the reflectance values (SLSTR).
# Imports
import os
# EOReader
from eoreader.reader import Reader
from eoreader.bands import YELLOW, Oa21, RED, SWIR_2, F1
# Declare the reader (only once)
reader = Reader()
# Create logger
import logging
from sertit import logs
logger = logging.getLogger("eoreader")
logs.init_logger(logger)
Sentinel-3 OLCI#
# First of all, let's focus on Sentinel-3 OLCI data
olci_path = os.path.join(
"/home", "data", "DATA", "PRODS", "S3",
"S3A_OL_1_EFR____20191215T105023_20191215T105323_20191216T153115_0179_052_322_2160_LN1_O_NT_002.zip"
)
olci_prod = reader.open(olci_path, remove_tmp=True)
olci_prod
eoreader.S3OlciProduct 'S3A_OL_1_EFR____20191215T105023_20191215T105323_20191216T153115_0179_052_322_2160_LN1_O_NT_002'
Attributes:
condensed_name: 20191215T105023_S3_OLCI_EFR
path: /home/data/DATA/PRODS/S3/S3A_OL_1_EFR____20191215T105023_20191215T105323_20191216T153115_0179_052_322_2160_LN1_O_NT_002.zip
constellation: Sentinel-3 OLCI
sensor type: Optical
product type: OL_1_EFR___
default pixel size: 300.0
default resolution: 300.0
acquisition datetime: 2019-12-15T10:50:23.000506
band mapping:
COASTAL_AEROSOL: 3
BLUE: 4
GREEN: 6
YELLOW: 7
RED: 8
VEGETATION_RED_EDGE_1: 11
VEGETATION_RED_EDGE_2: 12
VEGETATION_RED_EDGE_3: 16
NIR: 17
NARROW_NIR: 17
WATER_VAPOUR: 20
Oa01: 1
Oa02: 2
Oa09: 9
Oa10: 10
Oa13: 13
Oa14: 14
Oa15: 15
Oa18: 18
Oa19: 19
Oa21: 21
GREEN_I: 5
needs extraction: False
# Get the bands information
olci_prod.bands
eoreader.SpectralBand 'Oa01'
Attributes:
id: 1
eoreader_name: Oa01
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 400.0
Bandwidth (nm): 15.0
description: Aerosol correction, improved water constituent retrieval
eoreader.SpectralBand 'Oa02'
Attributes:
id: 2
eoreader_name: Oa02
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 412.5
Bandwidth (nm): 10.0
description: Yellow substance and detrital pigments (turbidity)
eoreader.SpectralBand 'Oa03'
Attributes:
id: 3
eoreader_name: COASTAL_AEROSOL
common_name: coastal
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 442.5
Bandwidth (nm): 10.0
description: Chlorophyll absorption maximum, biogeochemistry, vegetation
eoreader.SpectralBand 'Oa04'
Attributes:
id: 4
eoreader_name: BLUE
common_name: blue
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 490.0
Bandwidth (nm): 10.0
description: High Chlorophyll
eoreader.SpectralBand 'Oa05'
Attributes:
id: 5
eoreader_name: GREEN_I
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 510.0
Bandwidth (nm): 10.0
description: Chlorophyll, sediment, turbidity, red tide
eoreader.SpectralBand 'Oa06'
Attributes:
id: 6
eoreader_name: GREEN
common_name: green
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 560.0
Bandwidth (nm): 10.0
description: Chlorophyll reference (Chlorophyll minimum)
eoreader.SpectralBand 'Oa07'
Attributes:
id: 7
eoreader_name: YELLOW
common_name: yellow
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 620.0
Bandwidth (nm): 10.0
description: Sediment loading
eoreader.SpectralBand 'Oa08'
Attributes:
id: 8
eoreader_name: RED
common_name: red
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 665.0
Bandwidth (nm): 10.0
description: Chlorophyll (2nd Chlorophyll absorption maximum), sediment, yellow substance / vegetation
eoreader.SpectralBand 'Oa09'
Attributes:
id: 9
eoreader_name: Oa09
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 673.75
Bandwidth (nm): 7.5
description: For improved fluorescence retrieval and to better account for smile together with the bands 665 and 680 nm
eoreader.SpectralBand 'Oa10'
Attributes:
id: 10
eoreader_name: Oa10
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 681.25
Bandwidth (nm): 7.5
description: Chlorophyll fluorescence peak, red edge
eoreader.SpectralBand 'Oa11'
Attributes:
id: 11
eoreader_name: VEGETATION_RED_EDGE_1
common_name: rededge
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 708.75
Bandwidth (nm): 10.0
description: Chlorophyll fluorescence baseline, red edge transition
eoreader.SpectralBand 'Oa12'
Attributes:
id: 12
eoreader_name: VEGETATION_RED_EDGE_2
common_name: rededge
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 753.75
Bandwidth (nm): 7.5
description: O2 absorption/clouds, vegetation
eoreader.SpectralBand 'Oa13'
Attributes:
id: 13
eoreader_name: Oa13
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 761.25
Bandwidth (nm): 2.5
description: O2 absorption band/aerosol correction.
eoreader.SpectralBand 'Oa14'
Attributes:
id: 14
eoreader_name: Oa14
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 764.375
Bandwidth (nm): 3.75
description: Atmospheric correction
eoreader.SpectralBand 'Oa15'
Attributes:
id: 15
eoreader_name: Oa15
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 767.5
Bandwidth (nm): 2.5
description: O2A used for cloud top pressure, fluorescence over land
eoreader.SpectralBand 'Oa16'
Attributes:
id: 16
eoreader_name: VEGETATION_RED_EDGE_3
common_name: rededge
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 778.75
Bandwidth (nm): 15.0
description: Atmos. corr./aerosol corr.
eoreader.SpectralBand 'Oa17'
Attributes:
id: 17
eoreader_name: NIR
common_name: nir
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 865.0
Bandwidth (nm): 20.0
description: Atmospheric correction/aerosol correction, clouds, pixel co-registration
eoreader.SpectralBand 'Oa17'
Attributes:
id: 17
eoreader_name: NARROW_NIR
common_name: nir08
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 865.0
Bandwidth (nm): 20.0
description: Atmospheric correction/aerosol correction, clouds, pixel co-registration
eoreader.SpectralBand 'Oa18'
Attributes:
id: 18
eoreader_name: Oa18
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 885.0
Bandwidth (nm): 10.0
description: Water vapour absorption reference band. Common reference band with SLSTR instrument. Vegetation monitoring
eoreader.SpectralBand 'Oa19'
Attributes:
id: 19
eoreader_name: Oa19
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 900.0
Bandwidth (nm): 10.0
description: Water vapour absorption/vegetation monitoring (maximum reflectance)
eoreader.SpectralBand 'Oa20'
Attributes:
id: 20
eoreader_name: WATER_VAPOUR
common_name: nir09
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 940.0
Bandwidth (nm): 20.0
description: Water vapour absorption, Atmospheric correction/aerosol correction
eoreader.SpectralBand 'Oa21'
Attributes:
id: 21
eoreader_name: Oa21
common_name:
gsd (m): 300.0
asset_role: reflectance
Center wavelength (nm): 1020.0
Bandwidth (nm): 40.0
description: Atmospheric correction/aerosol correction
# Load the Yellow band and the far NIR one
# Please note that mapped band need to be called by their mapped name and the specific one with their true name
olci_bands = olci_prod.load([YELLOW, Oa21])
2023-11-02 14:47:45,156 - [DEBUG] - Loading bands ['YELLOW', 'Oa21']
2023-11-02 14:47:45,702 - [DEBUG] - Converting YELLOW to reflectance
/opt/conda/lib/python3.11/site-packages/rasterio/__init__.py:314: NotGeoreferencedWarning: The given matrix is equal to Affine.identity or its flipped counterpart. GDAL may ignore this matrix and save no geotransform without raising an error. This behavior is somewhat driver-specific.
dataset = writer(
2023-11-02 14:47:46,707 - [DEBUG] - Geocoding YELLOW
2023-11-02 14:47:55,454 - [DEBUG] - Converting Oa21 to reflectance
2023-11-02 14:47:56,452 - [DEBUG] - Geocoding Oa21
2023-11-02 14:48:05,389 - [DEBUG] - Read YELLOW
2023-11-02 14:48:05,434 - [DEBUG] - Manage nodata for band YELLOW
2023-11-02 14:48:05,437 - [DEBUG] - Converting YELLOW to reflectance
2023-11-02 14:48:06,391 - [DEBUG] - Read Oa21
2023-11-02 14:48:06,410 - [DEBUG] - Manage nodata for band Oa21
2023-11-02 14:48:06,414 - [DEBUG] - Converting Oa21 to reflectance
# Plot a subsampled version
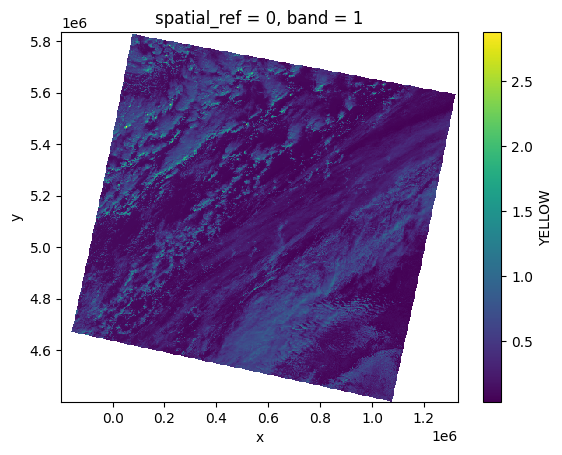
olci_bands[YELLOW][:, ::10, ::10].plot()
<matplotlib.collections.QuadMesh at 0x7f53f1927b10>
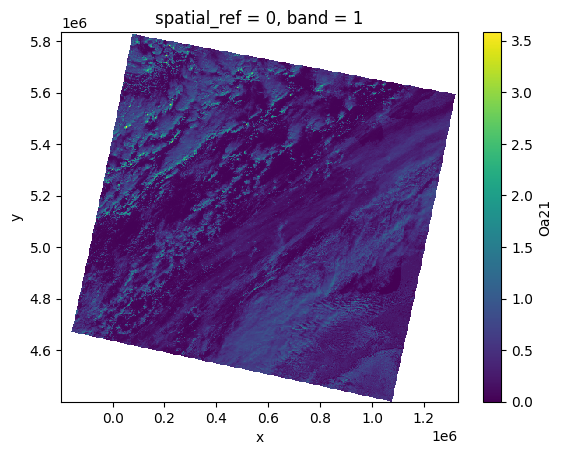
olci_bands[Oa21][:, ::10, ::10].plot()
<matplotlib.collections.QuadMesh at 0x7f53ec406b50>
Sentinel-3 SLSTR#
# Other SLSTR imports
from eoreader.keywords import SLSTR_VIEW, SLSTR_STRIPE, SLSTR_RAD_ADJUST
from eoreader.products import SlstrRadAdjustTuple, SlstrRadAdjust, SlstrView, SlstrStripe
# Then, let's focus on Sentinel-3 SLSTR data (extracted here, but a zip would work)
slstr_path = os.path.join(
"/home", "data", "DATA", "PRODS", "S3",
"S3B_SL_1_RBT____20191115T233722_20191115T234022_20191117T031722_0179_032_144_3420_LN2_O_NT_003.SEN3"
)
slstr_prod = reader.open(slstr_path, remove_tmp=True)
slstr_prod
eoreader.S3SlstrProduct 'S3B_SL_1_RBT____20191115T233722_20191115T234022_20191117T031722_0179_032_144_3420_LN2_O_NT_003'
Attributes:
condensed_name: 20191115T233722_S3_SLSTR_RBT
path: /home/data/DATA/PRODS/S3/S3B_SL_1_RBT____20191115T233722_20191115T234022_20191117T031722_0179_032_144_3420_LN2_O_NT_003.SEN3
constellation: Sentinel-3 SLSTR
sensor type: Optical
product type: SL_1_RBT___
default pixel size: 500.0
default resolution: 500.0
acquisition datetime: 2019-11-15T23:37:22.254773
band mapping:
GREEN: S1
RED: S2
NIR: S3
NARROW_NIR: S3
CIRRUS: S4
SWIR_1: S5
SWIR_2: S6
THERMAL_IR_1: S8
THERMAL_IR_2: S9
S7: S7
F1: F1
F2: F2
needs extraction: False
# Get the bands information
slstr_prod.bands
eoreader.SpectralBand 'F1'
Attributes:
id: F1
eoreader_name: F1
common_name:
gsd (m): 1000.0
asset_role: brightness_temperature
Center wavelength (nm): 3742.0
Bandwidth (nm): 398.0
description: Active fire, brightness temperature, 1km
eoreader.SpectralBand 'F2'
Attributes:
id: F2
eoreader_name: F2
common_name:
gsd (m): 1000.0
asset_role: brightness_temperature
Center wavelength (nm): 10854.0
Bandwidth (nm): 776.0
description: Active fire, brightness temperature, 1km
eoreader.SpectralBand 'S1'
Attributes:
id: S1
eoreader_name: GREEN
common_name: green
gsd (m): 500.0
asset_role: reflectance
Center wavelength (nm): 554.2700000000001
Bandwidth (nm): 19.259999999999998
description: Cloud screening, vegetation monitoring, aerosol
eoreader.SpectralBand 'S2'
Attributes:
id: S2
eoreader_name: RED
common_name: red
gsd (m): 500.0
asset_role: reflectance
Center wavelength (nm): 659.47
Bandwidth (nm): 19.25
description: NDVI, vegetation monitoring, aerosol
eoreader.SpectralBand 'S3'
Attributes:
id: S3
eoreader_name: NIR
common_name: nir
gsd (m): 500.0
asset_role: reflectance
Center wavelength (nm): 868.0
Bandwidth (nm): 20.6
description: NDVI, cloud flagging, pixel co-registration
eoreader.SpectralBand 'S3'
Attributes:
id: S3
eoreader_name: NARROW_NIR
common_name: nir08
gsd (m): 500.0
asset_role: reflectance
Center wavelength (nm): 868.0
Bandwidth (nm): 20.6
description: NDVI, cloud flagging, pixel co-registration
eoreader.SpectralBand 'S4'
Attributes:
id: S4
eoreader_name: CIRRUS
common_name: cirrus
gsd (m): 500.0
asset_role: reflectance
Center wavelength (nm): 1374.8
Bandwidth (nm): 20.8
description: Cirrus detection over land
eoreader.SpectralBand 'S5'
Attributes:
id: S5
eoreader_name: SWIR_1
common_name: swir16
gsd (m): 500.0
asset_role: reflectance
Center wavelength (nm): 1613.3999999999999
Bandwidth (nm): 60.68
description: Cloud clearing, ice, snow, vegetation monitoring
eoreader.SpectralBand 'S6'
Attributes:
id: S6
eoreader_name: SWIR_2
common_name: swir22
gsd (m): 1000.0
asset_role: reflectance
Center wavelength (nm): 2255.7
Bandwidth (nm): 50.15
description: Vegetation state and cloud clearing
eoreader.SpectralBand 'S7'
Attributes:
id: S7
eoreader_name: S7
common_name:
gsd (m): 1000.0
asset_role: brightness_temperature
Center wavelength (nm): 3742.0
Bandwidth (nm): 398.0
description: SST, LST, Active fire, brightness temperature, 1km
eoreader.SpectralBand 'S8'
Attributes:
id: S8
eoreader_name: THERMAL_IR_1
common_name: lwir11
gsd (m): 1000.0
asset_role: brightness_temperature
Center wavelength (nm): 10854.0
Bandwidth (nm): 776.0
description: SST, LST, Active fire, brightness temperature, 1km
eoreader.SpectralBand 'S9'
Attributes:
id: S9
eoreader_name: THERMAL_IR_2
common_name: lwir12
gsd (m): 1000.0
asset_role: brightness_temperature
Center wavelength (nm): 12022.5
Bandwidth (nm): 905.0
description: SST, LST, brightness temperature, 1km
# Same remark for mapped and specific band than above
# Not that native radiance band are converted into reflectance, whereas brilliance temperature bands are not
# Load bands with nadir view and stripe B
# (for bands that have a stripe B, the other will load their unique stripe, namely A or I)
# RED: only stripe A
# SWIR_2: has strip 1, B and TDI (c)
# F1: has only stripe I
slstr_bn_bands = slstr_prod.load([RED, SWIR_2, F1], slstr_view="n", slstr_stripe="b")
slstr_bn_bands_2 = slstr_prod.load([RED, SWIR_2, F1], **{SLSTR_VIEW: SlstrView.NADIR, SLSTR_STRIPE: SlstrStripe.B})
2023-11-02 14:48:10,152 - [DEBUG] - Loading bands ['RED', 'SWIR_2', 'F1']
2023-11-02 14:48:10,405 - [DEBUG] - Converting RED to reflectance
2023-11-02 14:48:16,239 - [DEBUG] - Geocoding RED
2023-11-02 14:48:22,045 - [DEBUG] - Converting SWIR_2 to reflectance
2023-11-02 14:48:27,513 - [DEBUG] - Geocoding SWIR_2
2023-11-02 14:48:32,910 - [DEBUG] - Geocoding F1
2023-11-02 14:48:37,485 - [DEBUG] - Read RED
2023-11-02 14:48:37,508 - [DEBUG] - Manage nodata for band RED
2023-11-02 14:48:37,512 - [DEBUG] - Converting RED to reflectance
2023-11-02 14:48:37,940 - [DEBUG] - Read SWIR_2
2023-11-02 14:48:37,960 - [DEBUG] - Manage nodata for band SWIR_2
2023-11-02 14:48:37,963 - [DEBUG] - Converting SWIR_2 to reflectance
2023-11-02 14:48:38,375 - [DEBUG] - Read F1
2023-11-02 14:48:38,396 - [DEBUG] - Manage nodata for band F1
2023-11-02 14:48:38,399 - [DEBUG] - Converting F1 to reflectance
2023-11-02 14:48:39,622 - [DEBUG] - Loading bands ['RED', 'SWIR_2', 'F1']
2023-11-02 14:48:39,623 - [DEBUG] - Read RED
2023-11-02 14:48:39,642 - [DEBUG] - Manage nodata for band RED
2023-11-02 14:48:39,646 - [DEBUG] - Converting RED to reflectance
2023-11-02 14:48:40,094 - [DEBUG] - Read SWIR_2
2023-11-02 14:48:40,113 - [DEBUG] - Manage nodata for band SWIR_2
2023-11-02 14:48:40,116 - [DEBUG] - Converting SWIR_2 to reflectance
2023-11-02 14:48:40,665 - [DEBUG] - Read F1
2023-11-02 14:48:40,688 - [DEBUG] - Manage nodata for band F1
2023-11-02 14:48:40,691 - [DEBUG] - Converting F1 to reflectance
# You can use the keywords by importing them or copy their value.
# Their values can be passed as strings or as an Enum
# However, it seems safer to import the keywords and use the enum
# The result should be the same
# Please bear in mind that oblique and nadir views are not stackable !
# However, you can stack different stripes
# (but you cannot load them at once and you should collocate them to be sure, their reprojection grid may vary as their GCP vary)
# Plot a subsampled version
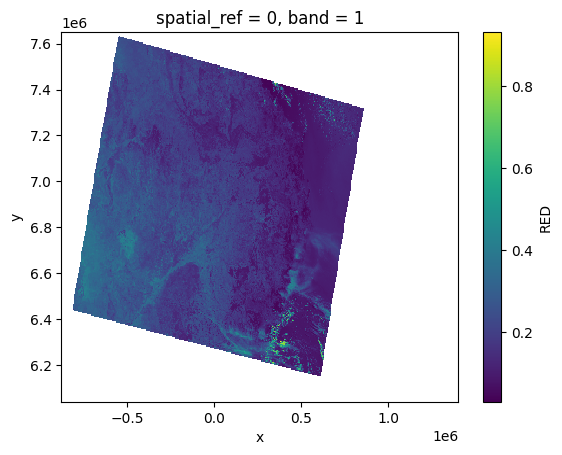
slstr_bn_bands[RED][:, ::10, ::10].plot()
<matplotlib.collections.QuadMesh at 0x7f53f62a6010>
slstr_bn_bands[SWIR_2][:, ::10, ::10].plot()
<matplotlib.collections.QuadMesh at 0x7f53a270d350>
slstr_bn_bands[F1][:, ::10, ::10].plot()
<matplotlib.collections.QuadMesh at 0x7f53a2661550>
# Sentinel-3 SLSTR radiance is not nominal, so EUMETSAT advises the user to make some radiance adjustments
# As stated here: https://www-cdn.eumetsat.int/files/2021-05/S3.PN-SLSTR-L1.08%20-%20i1r0%20-%20SLSTR%20L1%20PB%202.75-A%20and%201.53-B.pdf
# These coefficients have been added since the 06 version and several sets exist:
# The last one (S3.PN-SLSTR-L1.08, since 18/05/2021) which is also the default one
SlstrRadAdjust.S3_PN_SLSTR_L1_08
<SlstrRadAdjust.S3_PN_SLSTR_L1_08: SlstrRadAdjustTuple(S1_n=0.97, S2_n=0.98, S3_n=0.98, S4_n=1.0, S5_n=1.11, S6_n=1.13, S1_o=0.94, S2_o=0.95, S3_o=0.95, S4_o=1.0, S5_o=1.04, S6_o=1.07)>
# The two older sets given by EUMETSAT are the same
assert SlstrRadAdjust.S3_PN_SLSTR_L1_07 == SlstrRadAdjust.S3_PN_SLSTR_L1_06
SlstrRadAdjust.S3_PN_SLSTR_L1_07
<SlstrRadAdjust.S3_PN_SLSTR_L1_06: SlstrRadAdjustTuple(S1_n=1.0, S2_n=1.0, S3_n=1.0, S4_n=1.0, S5_n=1.12, S6_n=1.15, S1_o=1.0, S2_o=1.0, S3_o=1.0, S4_o=1.0, S5_o=1.2, S6_o=1.26)>
# Moreover, SNAP uses a different set with unknown origin (optional, in S3MPC Calibration)
SlstrRadAdjust.SNAP
<SlstrRadAdjust.SNAP: SlstrRadAdjustTuple(S1_n=1.0, S2_n=1.0, S3_n=1.0, S4_n=1.0, S5_n=1.12, S6_n=1.13, S1_o=1.0, S2_o=1.0, S3_o=1.0, S4_o=1.0, S5_o=1.15, S6_o=1.14)>
# A default set also exists, with every coefficient set to 1.0
SlstrRadAdjust.NONE
<SlstrRadAdjust.NONE: SlstrRadAdjustTuple(S1_n=1.0, S2_n=1.0, S3_n=1.0, S4_n=1.0, S5_n=1.0, S6_n=1.0, S1_o=1.0, S2_o=1.0, S3_o=1.0, S4_o=1.0, S5_o=1.0, S6_o=1.0)>
# You can use your own set by creating one.
# All the coefficients are set to 1.0 by default, so just modify the one you want
# The band keywords are {true_name}_{view_letter}
# RED is S2
user_set = SlstrRadAdjustTuple(S1_n=1.15, S2_o=1.12)
# However please bear in mind that if you want to reload the same band with a different adjustment,
# you have to remove the temporary process folder or the previous band will be reloaded.
slstr_prod.clean_tmp()
# To apply these sets when loading a band, just add the keyword when loading it
red_pn_08 = slstr_bn_bands[RED]
slstr_red_bn = slstr_prod.load(
RED,
**{
SLSTR_VIEW: SlstrView.NADIR,
SLSTR_STRIPE: SlstrStripe.B,
SLSTR_RAD_ADJUST: user_set
}
)
red_user = slstr_red_bn[RED]
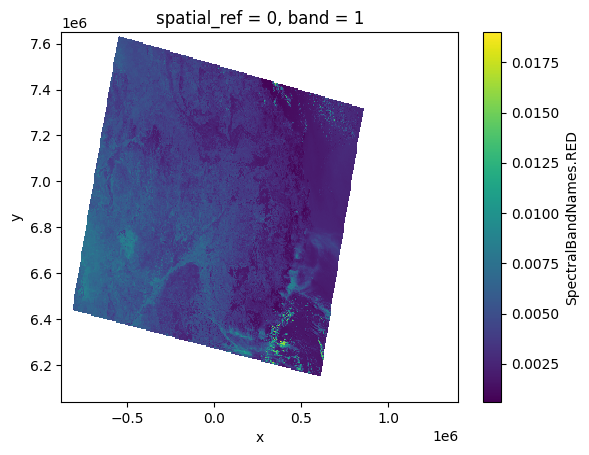
red_user[:, ::10, ::10].plot()
2023-11-02 14:48:42,997 - [DEBUG] - Loading bands ['RED']
2023-11-02 14:48:43,140 - [DEBUG] - Converting RED to reflectance
2023-11-02 14:48:48,554 - [DEBUG] - Geocoding RED
2023-11-02 14:48:53,472 - [DEBUG] - Read RED
2023-11-02 14:48:53,491 - [DEBUG] - Manage nodata for band RED
2023-11-02 14:48:53,494 - [DEBUG] - Converting RED to reflectance
<matplotlib.collections.QuadMesh at 0x7f53a26ebf90>
# We may need to collocate the bands if we want to work on two sets loaded apart
# Indeed, in EOReader, the bands are collocated when loaded together
# For example, if we wanted to work on the SWIR or F1 band,
# as we first loaded them with the RED, they are collocated to this band (the first one)
# Yet, their geodetic grid are different from the RED one (in and bn are slightly different than the an)
# So if we load on a second time only the SWIR or the F1 band, their are chances that the geocoding might be a little different
# The it is best to collocate the two bands just to be sure they will always match (and have the same size)
# To do so you could do:
from sertit import rasters
red_user = rasters.collocate(red_pn_08, other=red_user)
# Here, it is useless as we work on the master band
abs(red_pn_08 - red_user)[:, ::10, ::10].plot()
<matplotlib.collections.QuadMesh at 0x7f539db40850>