SAR example
Contents
SAR example#
Let’s use EOReader with SAR data.
Warning: SAR data is processed with SNAP, so be sure to have it installed and that GPT is in your path.
Imports#
import os
import logging
import matplotlib.pyplot as plt
# EOReader
from eoreader.reader import Reader
from eoreader.bands import VV, HH, VV_DSPK, HH_DSPK, HILLSHADE, SLOPE, to_str
from eoreader.env_vars import DEM_PATH
Create logger#
# Create logger
logger = logging.getLogger("eoreader")
logger.setLevel(logging.INFO)
# create console handler and set level to debug
ch = logging.StreamHandler()
ch.setLevel(logging.INFO)
# create formatter
formatter = logging.Formatter('%(message)s')
# add formatter to ch
ch.setFormatter(formatter)
# add ch to logger
logger.addHandler(ch)
Open the COSMO-SkyMed product#
Please be aware that:
EOReader will orthorectify your SAR data to get UTM tiles.
complex data is not handled as is, EOReader will convert them to ground range.
# First of all, we need some VHR data, let's use some COSMO-SkyMed data
path = os.path.join("/home", "data", "DATA", "PRODS", "COSMO", "1st_GEN", "1001512-735097")
# Open your product
prod = Reader().open(path, remove_tmp=True)
prod
eoreader.CskProduct 'CSKS4_DGM_B_HI_09_HH_RA_FF_20201008224018_20201008224025'
Attributes:
condensed_name: 20201008T224018_CSK_HI_DGM
path: /home/data/DATA/PRODS/COSMO/1st_GEN/1001512-735097
constellation: COSMO-SkyMed
sensor type: SAR
product type: DGM
default resolution: 5.0
acquisition datetime: 2020-10-08T22:40:18.446381
band mapping:
HH: HH
HH_DSPK: HH_DSPK
needs extraction: True
orbit direction: ASCENDING
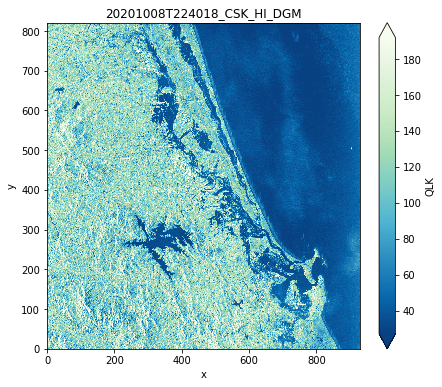
# Plot the quicklook
prod.plot()
/opt/conda/lib/python3.9/site-packages/rasterio/__init__.py:220: NotGeoreferencedWarning: Dataset has no geotransform, gcps, or rpcs. The identity matrix be returned.
s = DatasetReader(path, driver=driver, sharing=sharing, **kwargs)
/opt/conda/lib/python3.9/site-packages/rioxarray/_io.py:851: NotGeoreferencedWarning: Dataset has no geotransform, gcps, or rpcs. The identity matrix be returned.
warnings.warn(str(rio_warning.message), type(rio_warning.message)) # type: ignore
/opt/conda/lib/python3.9/site-packages/rasterio/__init__.py:230: NotGeoreferencedWarning: The given matrix is equal to Affine.identity or its flipped counterpart. GDAL may ignore this matrix and save no geotransform without raising an error. This behavior is somewhat driver-specific.
s = writer(path, mode, driver=driver,

# Get the band information
prod.bands
eoreader.SarBand 'HH'
Attributes:
id: HH
eoreader_name: HH
gsd (m): 5.0
asset_role: intensity
eoreader.SarBand 'HH_DSPK'
Attributes:
id: HH_DSPK
eoreader_name: HH_DSPK
gsd (m): 5.0
asset_role: intensity
# Print some data
print(f"Acquisition datetime: {prod.datetime}")
print(f"Condensed name: {prod.condensed_name}")
# Open here some more interesting geographical data: extent and footprint
extent = prod.extent()
footprint = prod.footprint()
base = extent.plot(color='cyan', edgecolor='black')
footprint.plot(ax=base, color='blue', edgecolor='black', alpha=0.5)
Acquisition datetime: 2020-10-08 22:40:18.446381
Condensed name: 20201008T224018_CSK_HI_DGM
Executing processing graph
.Copernicus_DSM_COG_10_N15_00_E108_00_DEM.tif
...10%....20%....30%....40%....50%....60%....70%....80%....90% done.
/opt/conda/lib/python3.9/site-packages/rasterio/__init__.py:220: NotGeoreferencedWarning: Dataset has no geotransform, gcps, or rpcs. The identity matrix be returned.
s = DatasetReader(path, driver=driver, sharing=sharing, **kwargs)
<AxesSubplot:>
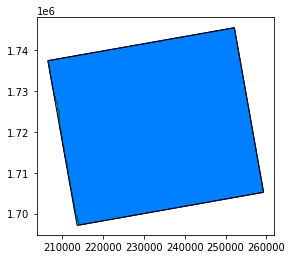
For SAR data, the footprint needs the orthorectified data !
For that, SNAP uses its own DEM, but you can change it when positionning the EOREADER_SNAP_DEM_NAME environment variable.
Available DEMs are:
ACE2_5MinACE30ASTER 1sec GDEMCopernicus 30m Global DEM(buggy for now, do not use it)Copernicus 90m Global DEM(buggy for now, do not use it)GETASSE30(by default)SRTM 1Sec HGTSRTM 3SecExternal DEM
Warning:
If External DEM is set, you must specify the DEM you want by positioning the EOREADER_DEM_PATH to a DEM that can be read by SNAP.
Load bands#
# Set the DEM
os.environ[DEM_PATH] = os.path.join("/home", "data", "DS2", "BASES_DE_DONNEES", "GLOBAL", "COPDEM_30m", "COPDEM_30m.vrt")
# Select some bands you wish to load without knowing if they exist
bands = [VV, HH, VV_DSPK, HH_DSPK, HILLSHADE, SLOPE]
# Only keep those selected
ok_bands = [band for band in bands if prod.has_band(band)]
# This product does not have VV band and HILLSHADE band cannot be computed from SAR band
print(to_str(ok_bands))
['HH', 'HH_DSPK', 'SLOPE']
# Load those bands as a dict of xarray.DataArray, with a 20m resolution
band_dict = prod.load(ok_bands, resolution=20.)
band_dict[HH]
Executing processing graph
first_line_time metadata value is null
last_line_time metadata value is null
...10%...21%...32%...43%...54%...65%...76%...87%. done.
<xarray.DataArray 'HH' (band: 1, y: 2474, x: 2689)>
array([[[nan, nan, nan, ..., nan, nan, nan],
[nan, nan, nan, ..., nan, nan, nan],
[nan, nan, nan, ..., nan, nan, nan],
...,
[nan, nan, nan, ..., nan, nan, nan],
[nan, nan, nan, ..., nan, nan, nan],
[nan, nan, nan, ..., nan, nan, nan]]], dtype=float32)
Coordinates:
* x (x) float64 2.058e+05 2.059e+05 ... 2.596e+05 2.596e+05
* y (y) float64 1.746e+06 1.746e+06 ... 1.697e+06 1.697e+06
* band (band) int64 1
spatial_ref int64 0
Attributes:
scale_factor: 1.0
add_offset: 0.0
long_name: HH
constellation: COSMO-SkyMed
constellation_id: CSK
product_path: /home/data/DATA/PRODS/COSMO/1st_GEN/1001512-735097
product_name: CSKS4_DGM_B_HI_09_HH_RA_FF_20201008224018_20201008224025
product_filename: 1001512-735097
instrument: SAR-2000
product_type: DGM
acquisition_date: 20201008T224018
condensed_name: 20201008T224018_CSK_HI_DGM
orbit_direction: ASCENDINGThis can lead the Terrain Correction step to create large nodata area when projecting on a DEM.
If it happens, you can set the keyword SAR_INTERP_NA to True when loading or stacking SAR data to fill these area with interpolated data.
from eoreader.keywords import SAR_INTERP_NA
band_dict = prod.load(
ok_bands,
resolution=20.,
**{SAR_INTERP_NA: True}
)
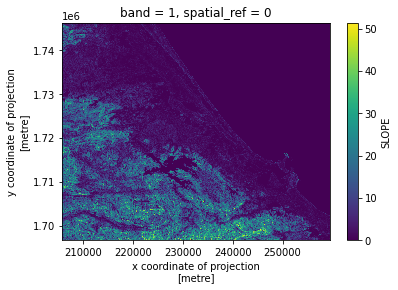
# Plot a subsampled version
band_dict[SLOPE][:, ::10, ::10].plot()
<matplotlib.collections.QuadMesh at 0x7fca4dadc7f0>

Stack some data#
# You can also stack those bands
stack = prod.stack(ok_bands)
stack
<xarray.DataArray 'HH_HH_DSPK_SLOPE' (z: 3, y: 9897, x: 10755)>
array([[[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...,
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan]],
[[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
...
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan],
[ nan, nan, nan, ..., nan,
nan, nan]],
[[ 0.4255329 , 0.4255329 , 0.4255329 , ..., 0. ,
0. , 0. ],
[ 0.4255329 , 0.4255329 , 0.4255329 , ..., 0. ,
0. , 0. ],
[ 0.4255329 , 0.4255329 , 0.4255329 , ..., 0. ,
0. , 0. ],
...,
[16.66219 , 16.66219 , 16.66219 , ..., 0.0870695 ,
0.0870695 , 0.0870695 ],
[16.66219 , 16.66219 , 16.66219 , ..., 0.0870695 ,
0.0870695 , 0.0870695 ],
[17.018255 , 17.018255 , 17.018255 , ..., 0.08737368,
0.08737368, 0.08737368]]], dtype=float32)
Coordinates:
spatial_ref int64 0
* x (x) float64 2.058e+05 2.058e+05 ... 2.596e+05 2.596e+05
* y (y) float64 1.746e+06 1.746e+06 ... 1.697e+06 1.697e+06
* z (z) MultiIndex
- variable (z) object 'HH' 'HH_DSPK' 'SLOPE'
- band (z) int64 1 1 1
Attributes:
long_name: HH HH_DSPK SLOPE
constellation: COSMO-SkyMed
constellation_id: CSK
product_path: /home/data/DATA/PRODS/COSMO/1st_GEN/1001512-735097
product_name: CSKS4_DGM_B_HI_09_HH_RA_FF_20201008224018_20201008224025
product_filename: 1001512-735097
instrument: SAR-2000
product_type: DGM
acquisition_date: 20201008T224018
condensed_name: 20201008T224018_CSK_HI_DGM
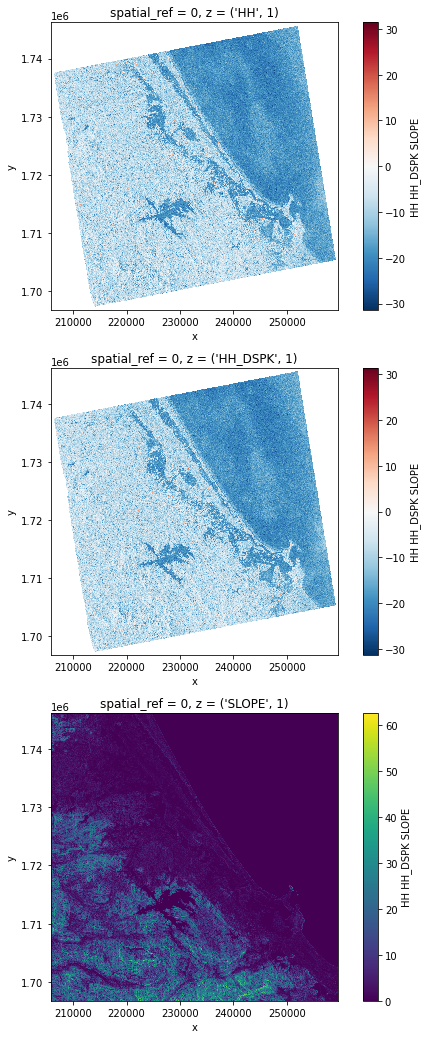
orbit_direction: ASCENDING# Plot a subsampled version
nrows = len(stack)
fig, axes = plt.subplots(nrows=nrows, figsize=(3 * nrows, 6 * nrows), subplot_kw={"box_aspect": 1})
for i in range(nrows):
stack[i, ::10, ::10].plot(x="x", y="y", ax=axes[i])

