Detect water with several constellations#
Let’s use EOReader for water detection, using several constellations over the same area.
Imports#
# Imports
import os
import numpy as np
import xarray as xr
from eoreader.reader import Reader
from eoreader.bands import NDWI
Core functions#
A function loading the Normalized Water Index (
NDWI)A function extracting water through a threshold
def load_ndwi(prod, pixel_size):
"""
Load NDWI index (and rename the array)
"""
# Read NDWI index
ndwi = prod.load(NDWI, pixel_size=pixel_size)[NDWI]
ndwi_name = f"{ndwi.attrs['constellation']} NDWI"
return ndwi.rename(ndwi_name)
def extract_water(ndwi):
"""
Extract water from NDWI index (and rename the array)
"""
# Assert water bodies when NDWI index > 0.2
water = xr.where(ndwi > 0.2, 1, 0)
# Set nodata where ndwi is nan.
# WARNING: the function xr.DataArray.where sets by default np.nan where the condition is false !
# See here: http://xarray.pydata.org/en/stable/generated/xarray.DataArray.where.html
water = water.where(~np.isnan(ndwi))
# Plot a subsampled version
water_name = f"{ndwi.attrs['constellation']} WATER"
water = water.rename(water_name)
water.attrs["long_name"] = "Water detection"
return water.rename(water_name)
Create logger#
# Create logger
import logging
from sertit import logs
logger = logging.getLogger("eoreader")
logs.init_logger(logger)
Linking some data#
Let’s take 3 products covering approximately the same area (over DAX in France):
One Landsat-8 OLI-TIRS collection 2
One Landsat-5 TM collection 2
One Sentinel-2 L1C
prod_folder = os.path.join("/home", "data", "DATA", "PRODS")
paths = [
# Landsat-8 OLI-TIRS collection 2
os.path.join(prod_folder, "LANDSATS_COL2", "LC08_L1TP_200030_20201220_20210310_02_T1.tar"),
# Landsat-5 TM collection 2
os.path.join(prod_folder, "LANDSATS_COL2", "LT05_L1TP_200030_20111110_20200820_02_T1.tar"),
# Sentinel-2 L2A
os.path.join(prod_folder, "S2", "PB 02.07+", "S2A_MSIL1C_20191215T110441_N0208_R094_T30TXP_20191215T114155.SAFE"),
]
Process these products#
# Create the reader
reader = Reader()
# Loop on all the products
water_arrays = []
ndwi_arrays = []
extents = []
for path in paths:
logger.info(f"*** {os.path.basename(path)} ***")
# Open the product
prod = reader.open(path, remove_tmp=True)
logger.info(prod)
# Get extents
extents.append(prod.extent())
# Read NDWI index
# Let's say we want a 60. meters pixel_size
ndwi = load_ndwi(prod, pixel_size=60.)
ndwi_arrays.append(ndwi)
# Extract water
water_arrays.append(extract_water(ndwi))
logger.info("\n")
2023-11-02 14:49:14,133 - [INFO] - *** LC08_L1TP_200030_20201220_20210310_02_T1.tar ***
2023-11-02 14:49:14,578 - [INFO] - eoreader.LandsatProduct 'LC08_L1TP_200030_20201220_20210310_02_T1'
Attributes:
condensed_name: 20201220T104856_L8_200030_OLI_TIRS
path: /home/data/DATA/PRODS/LANDSATS_COL2/LC08_L1TP_200030_20201220_20210310_02_T1.tar
constellation: Landsat-8
sensor type: Optical
product type: L1
default pixel size: 30.0
default resolution: 30.0
acquisition datetime: 2020-12-20T10:48:56
band mapping:
COASTAL_AEROSOL: 1
BLUE: 2
GREEN: 3
RED: 4
NIR: 5
NARROW_NIR: 5
CIRRUS: 9
SWIR_1: 6
SWIR_2: 7
THERMAL_IR_1: 10
THERMAL_IR_2: 11
PANCHROMATIC: 8
needs extraction: False
cloud cover: 16.36
tile name: 200030
2023-11-02 14:49:14,681 - [DEBUG] - Loading bands ['GREEN', 'NIR']
2023-11-02 14:49:14,700 - [DEBUG] - Read GREEN
2023-11-02 14:49:19,363 - [DEBUG] - Manage nodata for band GREEN
2023-11-02 14:49:20,326 - [DEBUG] - Converting GREEN to reflectance
2023-11-02 14:49:20,506 - [DEBUG] - Read NIR
2023-11-02 14:49:24,950 - [DEBUG] - Manage nodata for band NIR
2023-11-02 14:49:25,478 - [DEBUG] - Converting NIR to reflectance
2023-11-02 14:49:26,007 - [DEBUG] - Loading indices ['NDWI']
2023-11-02 14:49:26,284 - [INFO] -
2023-11-02 14:49:26,285 - [INFO] - *** LT05_L1TP_200030_20111110_20200820_02_T1.tar ***
2023-11-02 14:49:26,397 - [INFO] - eoreader.LandsatProduct 'LT05_L1TP_200030_20111110_20200820_02_T1'
Attributes:
condensed_name: 20111110T103612_L5_200030_TM
path: /home/data/DATA/PRODS/LANDSATS_COL2/LT05_L1TP_200030_20111110_20200820_02_T1.tar
constellation: Landsat-5
sensor type: Optical
product type: L1
default pixel size: 30.0
default resolution: 30.0
acquisition datetime: 2011-11-10T10:36:12
band mapping:
BLUE: 1
GREEN: 2
RED: 3
NIR: 4
NARROW_NIR: 4
SWIR_1: 5
SWIR_2: 7
THERMAL_IR_1: 6
THERMAL_IR_2: 6
needs extraction: False
cloud cover: 19.0
tile name: 200030
2023-11-02 14:49:26,474 - [DEBUG] - Loading bands ['GREEN', 'NIR']
2023-11-02 14:49:26,495 - [DEBUG] - Read GREEN
2023-11-02 14:49:29,421 - [DEBUG] - Manage nodata for band GREEN
2023-11-02 14:49:29,906 - [DEBUG] - Converting GREEN to reflectance
2023-11-02 14:49:30,044 - [DEBUG] - Read NIR
2023-11-02 14:49:33,321 - [DEBUG] - Manage nodata for band NIR
2023-11-02 14:49:33,868 - [DEBUG] - Converting NIR to reflectance
2023-11-02 14:49:34,297 - [DEBUG] - Loading indices ['NDWI']
2023-11-02 14:49:34,551 - [INFO] -
2023-11-02 14:49:34,552 - [INFO] - *** S2A_MSIL1C_20191215T110441_N0208_R094_T30TXP_20191215T114155.SAFE ***
2023-11-02 14:49:34,753 - [INFO] - eoreader.S2Product 'S2A_MSIL1C_20191215T110441_N0208_R094_T30TXP_20191215T114155'
Attributes:
condensed_name: 20191215T110441_S2_T30TXP_L1C_114155
path: /home/data/DATA/PRODS/S2/PB 02.07+/S2A_MSIL1C_20191215T110441_N0208_R094_T30TXP_20191215T114155.SAFE
constellation: Sentinel-2
sensor type: Optical
product type: MSIL1C
default pixel size: 10.0
default resolution: 10.0
acquisition datetime: 2019-12-15T11:04:41
band mapping:
COASTAL_AEROSOL: 01
BLUE: 02
GREEN: 03
RED: 04
VEGETATION_RED_EDGE_1: 05
VEGETATION_RED_EDGE_2: 06
VEGETATION_RED_EDGE_3: 07
NIR: 08
NARROW_NIR: 8A
WATER_VAPOUR: 09
CIRRUS: 10
SWIR_1: 11
SWIR_2: 12
needs extraction: False
cloud cover: 0.6668
tile name: T30TXP
2023-11-02 14:49:34,861 - [DEBUG] - Loading bands ['GREEN', 'NIR']
2023-11-02 14:49:34,936 - [DEBUG] - Read GREEN
2023-11-02 14:49:34,976 - [DEBUG] - Manage nodata for band GREEN
2023-11-02 14:49:35,249 - [DEBUG] - Converting GREEN to reflectance
2023-11-02 14:49:40,154 - [DEBUG] - Read NIR
2023-11-02 14:49:40,220 - [DEBUG] - Manage nodata for band NIR
2023-11-02 14:49:40,496 - [DEBUG] - Converting NIR to reflectance
2023-11-02 14:49:45,516 - [DEBUG] - Loading indices ['NDWI']
2023-11-02 14:49:45,962 - [INFO] -
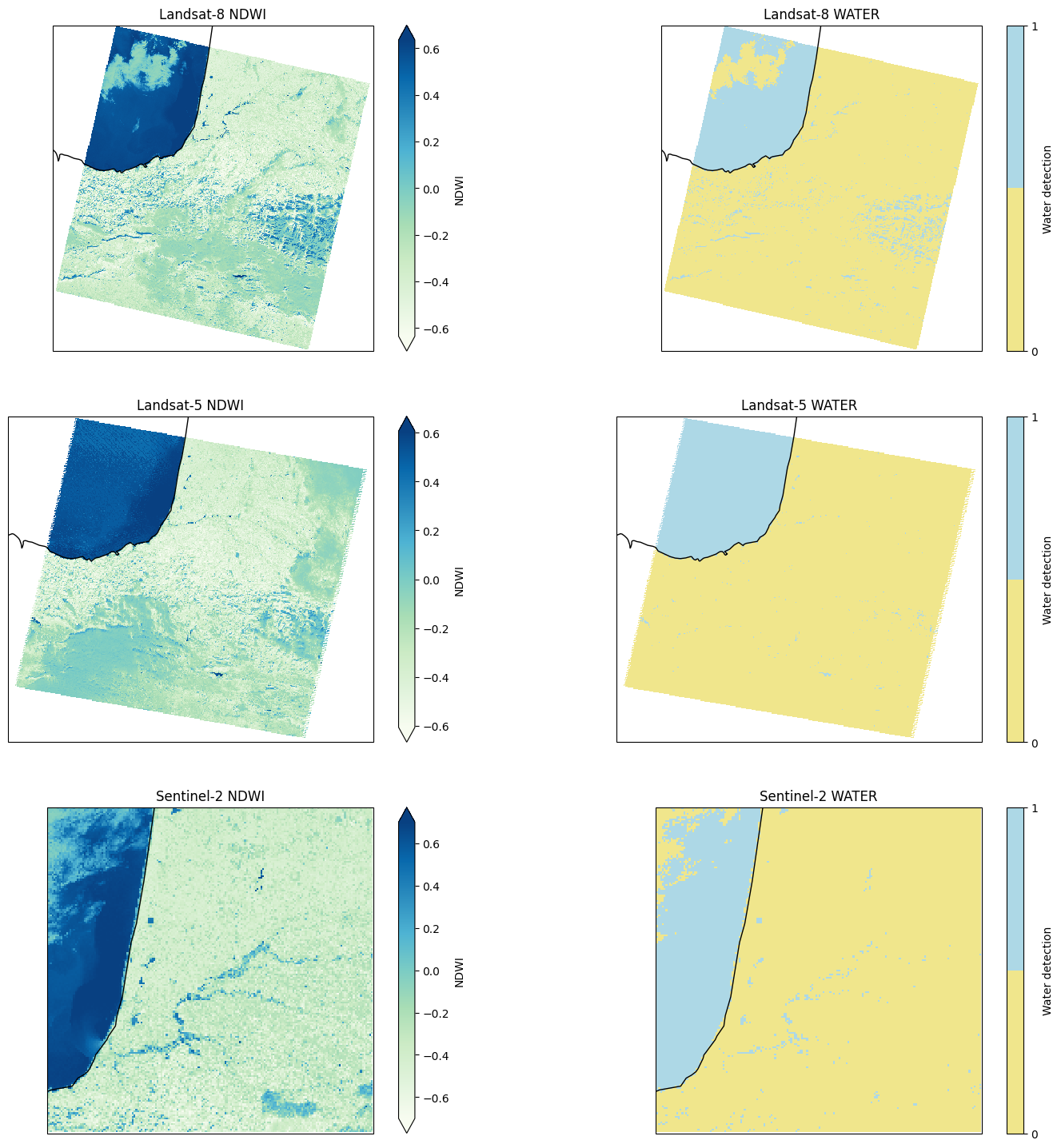
Plot the result#
# Plot the tiles
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
import cartopy.crs as ccrs
nrows = len(paths)
plt.figure(figsize=(6 * nrows, 6 * nrows))
cmap = ListedColormap(['khaki', 'lightblue'])
for i in range(nrows):
# Compute cartopy projection (EOReader always is in UTM)
extent = extents[i]
str_epsg = str(extent.crs.to_epsg())
proj = ccrs.UTM(
zone=str_epsg[-2:],
southern_hemisphere=str_epsg[2] == 7
)
# Get extent values
# The extents must be defined in the form (min_x, max_x, min_y, max_y)
bounds = extent.bounds
extent_val = [bounds.minx[0], bounds.maxx[0], bounds.miny[0], bounds.maxy[0]]
# Plot NDWI
axes = plt.subplot(nrows, 2, 2*i+1, projection=proj)
axes.set_extent(extent_val, proj)
ndwi_arrays[i][0, ::10, ::10].plot.imshow(origin='upper', extent=extent_val, transform=proj, cmap="GnBu", robust=True)
axes.coastlines(linewidth=1)
axes.set_title(ndwi_arrays[i].name)
# Plot water
axes = plt.subplot(nrows, 2, 2*i+2, projection=proj)
axes.set_extent(extent_val, proj)
water_arrays[i][0, ::10, ::10].plot.imshow(origin='upper', extent=extent_val, transform=proj, cmap=cmap, cbar_kwargs={'ticks': [0, 1]})
axes.coastlines(linewidth=1)
axes.set_title(water_arrays[i].name)
/opt/conda/lib/python3.11/site-packages/cartopy/io/__init__.py:241: DownloadWarning: Downloading: https://naturalearth.s3.amazonaws.com/10m_physical/ne_10m_coastline.zip
warnings.warn(f'Downloading: {url}', DownloadWarning)